ERA5 Data and Graphics Preparation for the Science-on-a-Sphere
This notebook will create all the necessary files and directories for a visualization on NOAA’s Science-on-a-Sphere. For this example, we will produce a map of 850 hPa geopotential heights, temperature, and horizontal wind, using ERA5, for a past weather event.
Overview:¶
Set up output directories for SoS
Specify the date/time range for your case
Use Cartopy’s
add_cyclic_pointmethod to avoid a blank seam at the datelineCreate small and large thumbnail graphics.
Create a standalone colorbar
Create a set of labels for each plot
Create 5 days worth of Science-on-a-Sphere-ready plots
Create an SoS playlist file
Prerequisites¶
| Concepts | Importance | Notes |
|---|---|---|
| Matplotlib | Necessary | |
| Cartopy | Necessary | |
| Xarray | Necessary | |
| Metpy | Necessary | |
| Linux command line / directory structure | Helpful |
Time to learn: 30 minutes
Imports¶
import xarray as xr
import numpy as np
import pandas as pd
import cartopy.feature as cfeature
import cartopy.crs as ccrs
import cartopy.util as cutil
from datetime import datetime as dt
import warnings
import metpy.calc as mpcalc
from metpy.units import units
import matplotlib.pyplot as pltDo not output warning messages
warnings.simplefilter("ignore")Set up output directories for SoS¶
The software that serves output for the Science-on-a-Sphere expects a directory structure as follows:
Top level directory: choose a name that is consistent with your case, e.g.: SS93
2nd level directory: choose a name that is consistent with the graphic, e.g.: SLP_500Z
3rd level directories:
2048: Contains the graphics (resolution: 2048x1024) that this notebook generates
labels: Contains one or two files:
(required) A text file,
labels.txt, which has as many lines as there are graphics in the 2048 file. Each line functions as the title of each graphic.(optional) A PNG file,
colorbar.png, which colorbar which will get overlaid on your map graphic.
media: Contains large and small thumbnails (
thumbnail_small.jpg,thumbnail_large.jpg) that serve as icons on the SoS iPad and SoS computer appsplaylist: A text file,
playlist.sos, which tells the SoS how to display your product
As an example, here is how the directory structure on our SoS looks for the products generated by this notebook. Our SoS computer stores locally-produced content in the /shared/sos/media/site-custom directory (note: the SoS directories are not network-accessible, so you won’t be able to cd into them). The ERA5 visualizations go in the ERA5 subfolder. Your top-level directory sits within the ERA5 folder.
sos@sos1:/shared/sos/media/site-custom/ERA5/cle_superbomb/SLP_500Z$ ls -R
.:
2048 labels media playlist
./2048:
ERA5_1978012600-fs8.png ERA5_1978012606-fs8.png ERA5_1978012612-fs8.png ERA5_1978012618-fs8.png
ERA5_1978012601-fs8.png ERA5_1978012607-fs8.png ERA5_1978012613-fs8.png ERA5_1978012619-fs8.png
ERA5_1978012602-fs8.png ERA5_1978012608-fs8.png ERA5_1978012614-fs8.png ERA5_1978012620-fs8.png
ERA5_1978012603-fs8.png ERA5_1978012609-fs8.png ERA5_1978012615-fs8.png ERA5_1978012621-fs8.png
ERA5_1978012604-fs8.png ERA5_1978012610-fs8.png ERA5_1978012616-fs8.png ERA5_1978012622-fs8.png
ERA5_1978012605-fs8.png ERA5_1978012611-fs8.png ERA5_1978012617-fs8.png ERA5_1978012623-fs8.png
./labels:
colorbar.png labels.txt
./media:
thumbnail_big.jpg thumbnail_small.jpg
./playlist:
playlist.sosDefine the 1st and 2nd-level directories.
# You define these
caseDir = 'Bliz2026'
prodDir = '850_T_Z_Wind'The 3rd-level directories follow from the 1st and 2nd.
# These remain as is
graphicsDir = caseDir + '/' + prodDir + '/2048/'
labelsDir = caseDir + '/' + prodDir + '/labels/'
mediaDir = caseDir + '/' + prodDir + '/media/'
playlistDir = caseDir + '/' + prodDir + '/playlist/'Create these directories via a Linux command
! mkdir -p {graphicsDir} {labelsDir} {mediaDir} {playlistDir}# Remove all PNGs in the graphics directory - COMMENT OUT if you don't want to do this!
! rm -f {graphicsDir}*.pngSpecify a starting and ending date/time, regional extent, vertical levels, and access/subset the ERA5¶
# Set the bounds of the map ... for the SoS, we will cover the entire globe.
## Do not change these values ##
latN = 90
latS = -90
lonW = 0
lonE = 360
latRange = np.arange(latS, latN + 0.1,.5) # ensure +90 is included
lonRange = np.arange(lonW, lonE, .5)
##
# Select the map projection for your figure.
proj_map = ccrs.PlateCarree() # For SoS maps
# Select the projection on which the dataset is based
proj_data = ccrs.PlateCarree()
# Select the resolution of the Cartopy cartographic features: coarsest Natural Earth, and no counties
res = '110m' # Least detailed, best for large/global maps
# Specify desired pressure levels: in this case, a list of one or more
plevel_list = [850]
startYear = 2026
startMonth = 2
startDay = 20
startHour = 0
startMinute = 0
startDateTime = dt(startYear,startMonth,startDay, startHour, startMinute)
endYear = 2026
endMonth = 2
endDay = 24
endHour = 18
endMinute = 0
endDateTime = dt(endYear,endMonth,endDay, endHour, endMinute)
delta_time = endDateTime - startDateTime
time_range_max = 7*86400
if (delta_time.total_seconds() > time_range_max):
raise RuntimeError("Your time range must not exceed 7 days. Go back and try again.")
# Create a list of date and times based on what we specified for the initial and final times,
# using Pandas’ date_range function
dateRange = pd.date_range(startDateTime, endDateTime,freq="6h")Access either the cloud-served or local ERA5 repository. Then subset it according to your choices above.¶
endDate = dt(2023,1,10)
if (endDateTime <= endDate): # Use WeatherBench archive
cloud_source = True
ds = xr.open_dataset(
'gs://weatherbench2/datasets/era5/1959-2023_01_10-wb13-6h-1440x721.zarr',
chunks={'time': 48},
consolidated=True,
engine='zarr'
)
# Attach units to the pressure coordinate
ds.coords['level'].attrs['units'] = 'hPa'
# Rename the variable names in this dataset so they use their corresponding short_name attributes
# Construct a dictionary whose keys are the original variable names, and whose values are their
# short names
rename_data_vars = {}
seen_short_names = set()
for var_name in ds.data_vars:
short = ds[var_name].attrs.get("short_name")
# Only rename if:
# 1. short_name exists
# 2. We haven't already used it
if short and short not in seen_short_names:
rename_data_vars[var_name] = short
seen_short_names.add(short)
# Apply renaming
ds = ds.rename(rename_data_vars)
else: # Use local archive
import glob, os
cloud_source = False
input_directory = '/free/ktyle/era5'
files = glob.glob(os.path.join(input_directory,'*_era5.nc'))
ds = xr.open_mfdataset(files,coords='minimal',compat='override')
# Rename two of the coordinate variables so they match what is in the WeatherBench archive
ds = ds.rename({'valid_time': 'time', 'pressure_level': 'level'})
# Perform the initial temporal and vertical level subset
ds = ds.sel(time = dateRange, longitude = lonRange, latitude = latRange, level = plevel_list)Examine the subsetted Dataset¶
Expand the Data variables section to see their names and dimensions!
dsTo avoid contouring problems across the dateline, create a function so longitudes run from -180 to 180 instead of 0 to 360.¶
def reorder_longitudes(da):
"""
This function reorders the longitudes in a global grid from 0 --> 360 to -180 --> 180
"""
da = da.assign_coords(longitude=(((da.longitude + 180 ) % 360 ) - 180))
return da.sortby(da['longitude'])Create DataArray objects for the data variables we are interested in. Reorder their longitude dimension so they begin at 180 instead of 0.¶
q, t, u, v, w, and z are all four-dimensional; i.e., they also include isobaric levels. The others (e.g. msl and sst) are all three-dimensional; i.e., they do not have a vertical dimension.
# Select only those variables that you need for your product. Comment out or remove those you don't need.
## 4-D variables
T = reorder_longitudes(ds['t'])
U = reorder_longitudes(ds['u'])
V = reorder_longitudes(ds['v'])
Z = reorder_longitudes(ds['z'])
# 3-D variables
#SLP = reorder_longitudes(ds['msl'])For the 4-D variables, select just a single isobaric surface from our initial list. Load in the data into RAM via the compute function.¶
This step, involving actual data retrieval, may take some time, particularly if you are retrieving from the cloud-served WeatherBench Zarr archive.
For the cloud-served archive, each variable will take approximately 10-20 seconds normally.
For the cloud-served archive, each variable will take approximately 5-10 seconds normally. If the servers are busy, it will take longer. Be patient!
p_level = 850
# 4-d variables; as above, comment out or delete those you don't need
%time Tp = T.sel(level=p_level).compute()
%time Up = U.sel(level=p_level).compute()
%time Vp = V.sel(level=p_level).compute()
%time Zp = Z.sel(level=p_level).compute()
# 3-d variables; similarly, comment out or delete those you don't need
#%time SLPp = SLP.compute()CPU times: user 7.2 s, sys: 1.62 s, total: 8.82 s
Wall time: 5.58 s
CPU times: user 8.22 s, sys: 910 ms, total: 9.13 s
Wall time: 6.05 s
CPU times: user 8.66 s, sys: 774 ms, total: 9.43 s
Wall time: 6.35 s
CPU times: user 7.2 s, sys: 636 ms, total: 7.83 s
Wall time: 4.83 s
%time magic directiveThis Jupyter lab directive will display how long each line beginning with the directive takes to execute.
Create objects for the relevant coordinate arrays; in this case, longitude, latitude, and time.¶
Remember to use one of the variables that you selected above ... in this example, we’ll use temperature on the selected isobaric surface (Tp)
lons, lats, times= Tp.longitude, Tp.latitude, Tp.timelonsPerform any necessary diagnostics, rescaling, smoothing and/or unit conversions.¶
For this example, we only need to convert geopotential to geopotential height diagnostic-wise. Rescaling and smoothing are not necessary. We will do some unit conversions.
Zdam = mpcalc.geopotential_to_height(Zp).metpy.convert_units('dam')
Ukts = Up.metpy.convert_units('kts')
Vkts = Vp.metpy.convert_units('kts')
Tc = Tp.metpy.convert_units('degC')Set contour levels for temperature, after first inspecting its range of values.
TcTc.min().values,Tc.max().values(array(-43.74002, dtype=float32), array(32.545868, dtype=float32))tContours = np.arange(-45,39,3)For the purposes of plotting geopotential heights in decameters, choose an appropriate contour interval and range of values ... for geopotential heights, a common convention is: from surface up through 700 hPa: 3 dam; above that, 6 dam to 400 and then 9 or 12 dam from 400 and above.
if (p_level >= 700):
incZ = 3
elif (p_level >= 400):
incZ = 6
elif (p_level >= 150):
incZ = 9
else:
incZ = 12
zContours = np.arange(0,3000,incZ)Create a plot of temperature, wind, and geopotential height.¶
Let’s create a plot for the first time in the time range (isel=0), just to ensure all looks good.
The Science on a Sphere treats titles and colorbars as separate layers. Thus, in these next cells we will not generate nor include them in the figure.
Additionally, the sphere expects its graphics to have a resolution of 2048x1024.
Instead of restricting the map region via
set_extent, we useset_global, to direct Matplotlib to use the full (global) x- and y- dimension ranges.Finally, by default, Matplotlib includes a border frame around each
Axes. We don’t want that included on the sphere-projected graphic either.
T0 = Tc.isel(time=0)
U0 = Ukts.isel(time=0)
V0 = Vkts.isel(time=0)
Z0 = Zdam.isel(time=0)Z0Set colors for map backgrounds and wind barbs¶
coast_border_col = 'dimgrey'
state_col = 'lightgrey'
wind_col = 'whitesmoke'dpi = 100
fig = plt.figure(figsize=(2048/dpi, 1024/dpi))
ax = fig.add_subplot(1,1,1,projection=ccrs.PlateCarree(central_longitude=180), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.BORDERS.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.STATES.with_scale(res), edgecolor=state_col)
# Temperature (T) contour fills
# Note we don't need the transform argument since the map/data projections are the same, but we'll leave it in
CF = ax.contourf(lons,lats,T0,levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
# Height (Z) contour lines
CL = ax.contour(lons,lats,Z0,zContours,linewidths=1.25,colors='yellow', transform=proj_data)
ax.clabel(CL, inline_spacing=0.2, fontsize=8, fmt='%.0f')
fig.tight_layout(pad=.01)
# Plotting wind barbs uses the ax.barbs method. Here, you can't pass in the DataArray directly; you can only pass in the array's values.
# Also need to sample (skip) a selected # of points to keep the plot readable.
skip = 8
# Default length of wind barbs is 7. Shorten them a bit.
ax.barbs(lons[::skip],lats[::skip],U0[::skip,::skip].values, V0[::skip,::skip].values, color=wind_col, length = 5, transform=proj_data);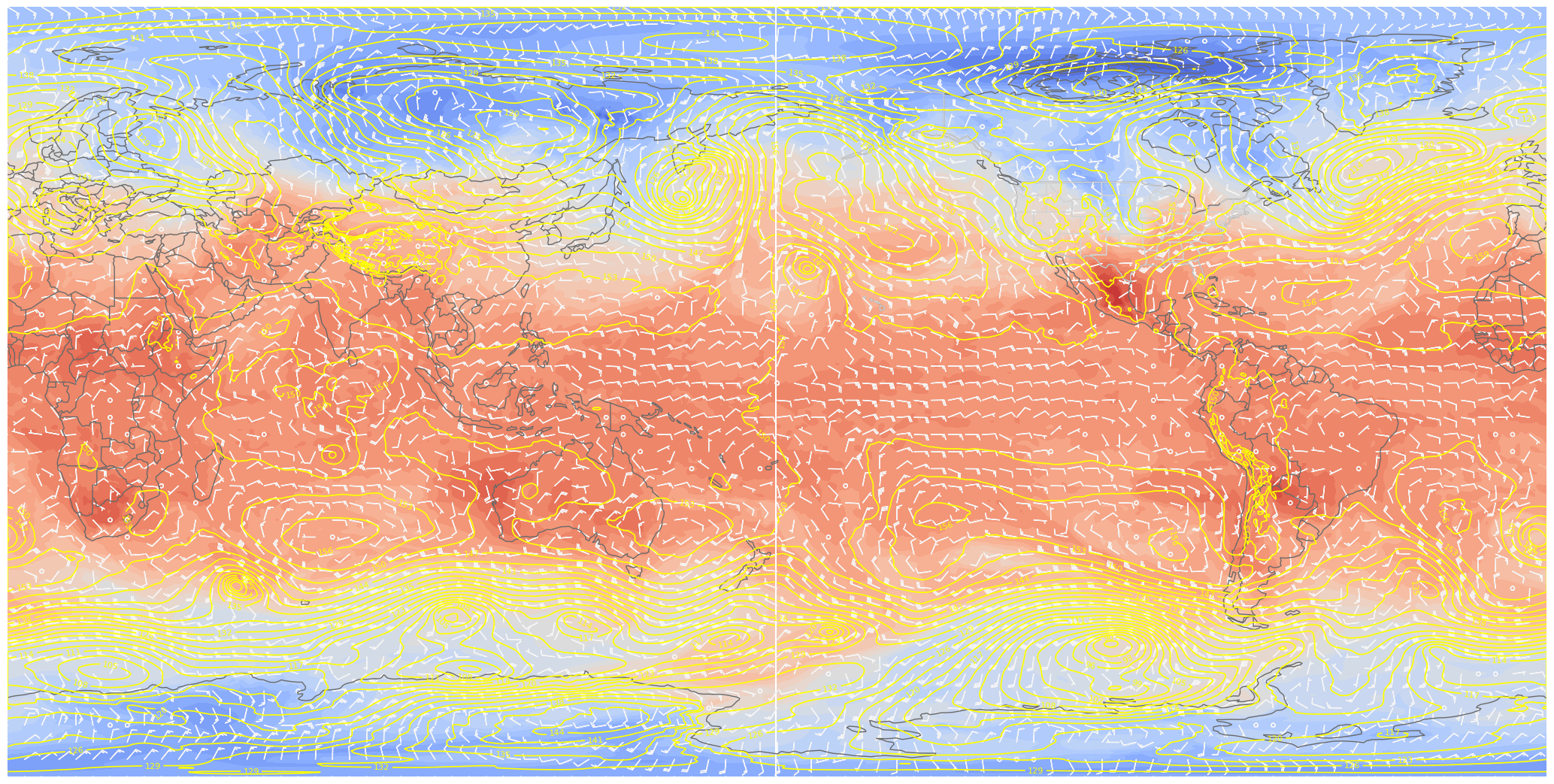
Hmmm, notice something? Let’s now try plotting on an Orthographic projection, somewhat akin to how it would look on the sphere.
dpi = 100
fig = plt.figure(figsize=(2048/dpi, 1024/dpi))
ax = fig.add_subplot(1,1,1,projection=ccrs.Orthographic(central_longitude=180), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.BORDERS.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.STATES.with_scale(res), edgecolor=state_col)
# Temperature (T) contour fills
# Note we don't need the transform argument since the map/data projections are the same, but we'll leave it in
CF = ax.contourf(lons,lats,T0,levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
# Height (Z) contour lines
CL = ax.contour(lons,lats,Z0,zContours,linewidths=1.25,colors='yellow', transform=proj_data)
ax.clabel(CL, inline_spacing=0.2, fontsize=8, fmt='%.0f')
fig.tight_layout(pad=.01)
# Plotting wind barbs uses the ax.barbs method. Here, you can't pass in the DataArray directly; you can only pass in the array's values.
# Also need to sample (skip) a selected # of points to keep the plot readable.
skip = 8
# Default length of wind barbs is 7. Shorten them a bit.
ax.barbs(lons[::skip],lats[::skip],U0[::skip,::skip].values, V0[::skip,::skip].values, color=wind_col, length = 5, transform=proj_data);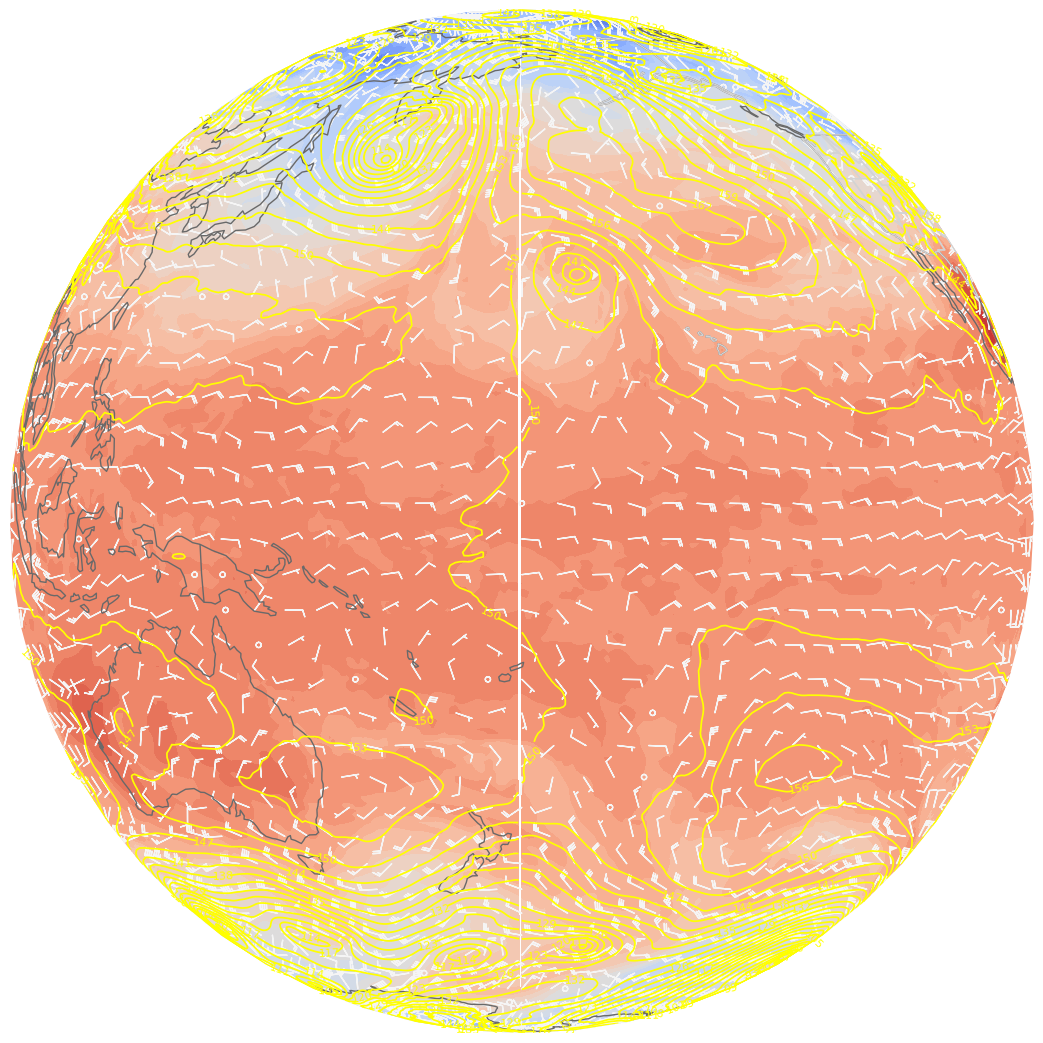
There “seams” to be a problem close to the dateline. This is because the longitude ends at 179.5 E ... leaving a gap between 179.5 and 180 degrees. Fortunately, Cartopy includes the add_cyclic_point method in its utilities class to deal with this rather common phenomenon in global gridded datasets.
# add cyclic points to data and longitudes
# latitudes are unchanged in 1-dimension
Z_cyc, lons_cyc = cutil.add_cyclic_point(Z0, lons)
T_cyc, lons_cyc = cutil.add_cyclic_point(T0, lons)
U_cyc, lons_cyc = cutil.add_cyclic_point(U0, lons)
V_cyc, lons_cyc = cutil.add_cyclic_point(V0, lons)Z_cycmasked_array(
data=[[130.20805, 130.20805, 130.20805, ..., 130.20805, 130.20805,
130.20805],
[130.12645, 130.12645, 130.12389, ..., 130.12898, 130.12645,
130.12645],
[130.04483, 130.03972, 130.03717, ..., 130.05757, 130.04993,
130.04483],
...,
[133.61276, 133.60257, 133.59236, ..., 133.63062, 133.62296,
133.61276],
[132.88081, 132.87572, 132.8706 , ..., 132.88846, 132.88591,
132.88081],
[131.87088, 131.87088, 131.87088, ..., 131.87088, 131.87088,
131.87088]],
mask=False,
fill_value=np.float64(1e+20),
dtype=float32)lons_cycmasked_array(data=[-180. , -179.5, -179. , -178.5, -178. , -177.5, -177. ,
-176.5, -176. , -175.5, -175. , -174.5, -174. , -173.5,
-173. , -172.5, -172. , -171.5, -171. , -170.5, -170. ,
-169.5, -169. , -168.5, -168. , -167.5, -167. , -166.5,
-166. , -165.5, -165. , -164.5, -164. , -163.5, -163. ,
-162.5, -162. , -161.5, -161. , -160.5, -160. , -159.5,
-159. , -158.5, -158. , -157.5, -157. , -156.5, -156. ,
-155.5, -155. , -154.5, -154. , -153.5, -153. , -152.5,
-152. , -151.5, -151. , -150.5, -150. , -149.5, -149. ,
-148.5, -148. , -147.5, -147. , -146.5, -146. , -145.5,
-145. , -144.5, -144. , -143.5, -143. , -142.5, -142. ,
-141.5, -141. , -140.5, -140. , -139.5, -139. , -138.5,
-138. , -137.5, -137. , -136.5, -136. , -135.5, -135. ,
-134.5, -134. , -133.5, -133. , -132.5, -132. , -131.5,
-131. , -130.5, -130. , -129.5, -129. , -128.5, -128. ,
-127.5, -127. , -126.5, -126. , -125.5, -125. , -124.5,
-124. , -123.5, -123. , -122.5, -122. , -121.5, -121. ,
-120.5, -120. , -119.5, -119. , -118.5, -118. , -117.5,
-117. , -116.5, -116. , -115.5, -115. , -114.5, -114. ,
-113.5, -113. , -112.5, -112. , -111.5, -111. , -110.5,
-110. , -109.5, -109. , -108.5, -108. , -107.5, -107. ,
-106.5, -106. , -105.5, -105. , -104.5, -104. , -103.5,
-103. , -102.5, -102. , -101.5, -101. , -100.5, -100. ,
-99.5, -99. , -98.5, -98. , -97.5, -97. , -96.5,
-96. , -95.5, -95. , -94.5, -94. , -93.5, -93. ,
-92.5, -92. , -91.5, -91. , -90.5, -90. , -89.5,
-89. , -88.5, -88. , -87.5, -87. , -86.5, -86. ,
-85.5, -85. , -84.5, -84. , -83.5, -83. , -82.5,
-82. , -81.5, -81. , -80.5, -80. , -79.5, -79. ,
-78.5, -78. , -77.5, -77. , -76.5, -76. , -75.5,
-75. , -74.5, -74. , -73.5, -73. , -72.5, -72. ,
-71.5, -71. , -70.5, -70. , -69.5, -69. , -68.5,
-68. , -67.5, -67. , -66.5, -66. , -65.5, -65. ,
-64.5, -64. , -63.5, -63. , -62.5, -62. , -61.5,
-61. , -60.5, -60. , -59.5, -59. , -58.5, -58. ,
-57.5, -57. , -56.5, -56. , -55.5, -55. , -54.5,
-54. , -53.5, -53. , -52.5, -52. , -51.5, -51. ,
-50.5, -50. , -49.5, -49. , -48.5, -48. , -47.5,
-47. , -46.5, -46. , -45.5, -45. , -44.5, -44. ,
-43.5, -43. , -42.5, -42. , -41.5, -41. , -40.5,
-40. , -39.5, -39. , -38.5, -38. , -37.5, -37. ,
-36.5, -36. , -35.5, -35. , -34.5, -34. , -33.5,
-33. , -32.5, -32. , -31.5, -31. , -30.5, -30. ,
-29.5, -29. , -28.5, -28. , -27.5, -27. , -26.5,
-26. , -25.5, -25. , -24.5, -24. , -23.5, -23. ,
-22.5, -22. , -21.5, -21. , -20.5, -20. , -19.5,
-19. , -18.5, -18. , -17.5, -17. , -16.5, -16. ,
-15.5, -15. , -14.5, -14. , -13.5, -13. , -12.5,
-12. , -11.5, -11. , -10.5, -10. , -9.5, -9. ,
-8.5, -8. , -7.5, -7. , -6.5, -6. , -5.5,
-5. , -4.5, -4. , -3.5, -3. , -2.5, -2. ,
-1.5, -1. , -0.5, 0. , 0.5, 1. , 1.5,
2. , 2.5, 3. , 3.5, 4. , 4.5, 5. ,
5.5, 6. , 6.5, 7. , 7.5, 8. , 8.5,
9. , 9.5, 10. , 10.5, 11. , 11.5, 12. ,
12.5, 13. , 13.5, 14. , 14.5, 15. , 15.5,
16. , 16.5, 17. , 17.5, 18. , 18.5, 19. ,
19.5, 20. , 20.5, 21. , 21.5, 22. , 22.5,
23. , 23.5, 24. , 24.5, 25. , 25.5, 26. ,
26.5, 27. , 27.5, 28. , 28.5, 29. , 29.5,
30. , 30.5, 31. , 31.5, 32. , 32.5, 33. ,
33.5, 34. , 34.5, 35. , 35.5, 36. , 36.5,
37. , 37.5, 38. , 38.5, 39. , 39.5, 40. ,
40.5, 41. , 41.5, 42. , 42.5, 43. , 43.5,
44. , 44.5, 45. , 45.5, 46. , 46.5, 47. ,
47.5, 48. , 48.5, 49. , 49.5, 50. , 50.5,
51. , 51.5, 52. , 52.5, 53. , 53.5, 54. ,
54.5, 55. , 55.5, 56. , 56.5, 57. , 57.5,
58. , 58.5, 59. , 59.5, 60. , 60.5, 61. ,
61.5, 62. , 62.5, 63. , 63.5, 64. , 64.5,
65. , 65.5, 66. , 66.5, 67. , 67.5, 68. ,
68.5, 69. , 69.5, 70. , 70.5, 71. , 71.5,
72. , 72.5, 73. , 73.5, 74. , 74.5, 75. ,
75.5, 76. , 76.5, 77. , 77.5, 78. , 78.5,
79. , 79.5, 80. , 80.5, 81. , 81.5, 82. ,
82.5, 83. , 83.5, 84. , 84.5, 85. , 85.5,
86. , 86.5, 87. , 87.5, 88. , 88.5, 89. ,
89.5, 90. , 90.5, 91. , 91.5, 92. , 92.5,
93. , 93.5, 94. , 94.5, 95. , 95.5, 96. ,
96.5, 97. , 97.5, 98. , 98.5, 99. , 99.5,
100. , 100.5, 101. , 101.5, 102. , 102.5, 103. ,
103.5, 104. , 104.5, 105. , 105.5, 106. , 106.5,
107. , 107.5, 108. , 108.5, 109. , 109.5, 110. ,
110.5, 111. , 111.5, 112. , 112.5, 113. , 113.5,
114. , 114.5, 115. , 115.5, 116. , 116.5, 117. ,
117.5, 118. , 118.5, 119. , 119.5, 120. , 120.5,
121. , 121.5, 122. , 122.5, 123. , 123.5, 124. ,
124.5, 125. , 125.5, 126. , 126.5, 127. , 127.5,
128. , 128.5, 129. , 129.5, 130. , 130.5, 131. ,
131.5, 132. , 132.5, 133. , 133.5, 134. , 134.5,
135. , 135.5, 136. , 136.5, 137. , 137.5, 138. ,
138.5, 139. , 139.5, 140. , 140.5, 141. , 141.5,
142. , 142.5, 143. , 143.5, 144. , 144.5, 145. ,
145.5, 146. , 146.5, 147. , 147.5, 148. , 148.5,
149. , 149.5, 150. , 150.5, 151. , 151.5, 152. ,
152.5, 153. , 153.5, 154. , 154.5, 155. , 155.5,
156. , 156.5, 157. , 157.5, 158. , 158.5, 159. ,
159.5, 160. , 160.5, 161. , 161.5, 162. , 162.5,
163. , 163.5, 164. , 164.5, 165. , 165.5, 166. ,
166.5, 167. , 167.5, 168. , 168.5, 169. , 169.5,
170. , 170.5, 171. , 171.5, 172. , 172.5, 173. ,
173.5, 174. , 174.5, 175. , 175.5, 176. , 176.5,
177. , 177.5, 178. , 178.5, 179. , 179.5, 180. ],
mask=False,
fill_value=1e+20)dpi = 100
fig = plt.figure(figsize=(2048/dpi, 1024/dpi))
ax = fig.add_subplot(1,1,1, projection=ccrs.Orthographic(central_longitude=180), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.BORDERS.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.STATES.with_scale(res), edgecolor=state_col)
# Temperature (T) contour fills
# Note we don't need the transform argument since the map/data projections are the same, but we'll leave it in
CF = ax.contourf(lons_cyc,lats,T_cyc,levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
# Height (Z) contour lines
CL = ax.contour(lons_cyc,lats,Z_cyc,zContours,linewidths=1.25,colors='yellow', transform=proj_data)
ax.clabel(CL, inline_spacing=0.2, fontsize=8, fmt='%.0f')
fig.tight_layout(pad=.01)
# Plotting wind barbs uses the ax.barbs method. Here, you can't pass in the DataArray directly; you can only pass in the array's values.
# Also need to sample (skip) a selected # of points to keep the plot readable.
skip = 8
# Default length of wind barbs is 7. Shorten them a bit.
ax.barbs(lons_cyc[::skip],lats[::skip],U_cyc[::skip,::skip], V_cyc[::skip,::skip], color=wind_col, length = 5, transform=proj_data);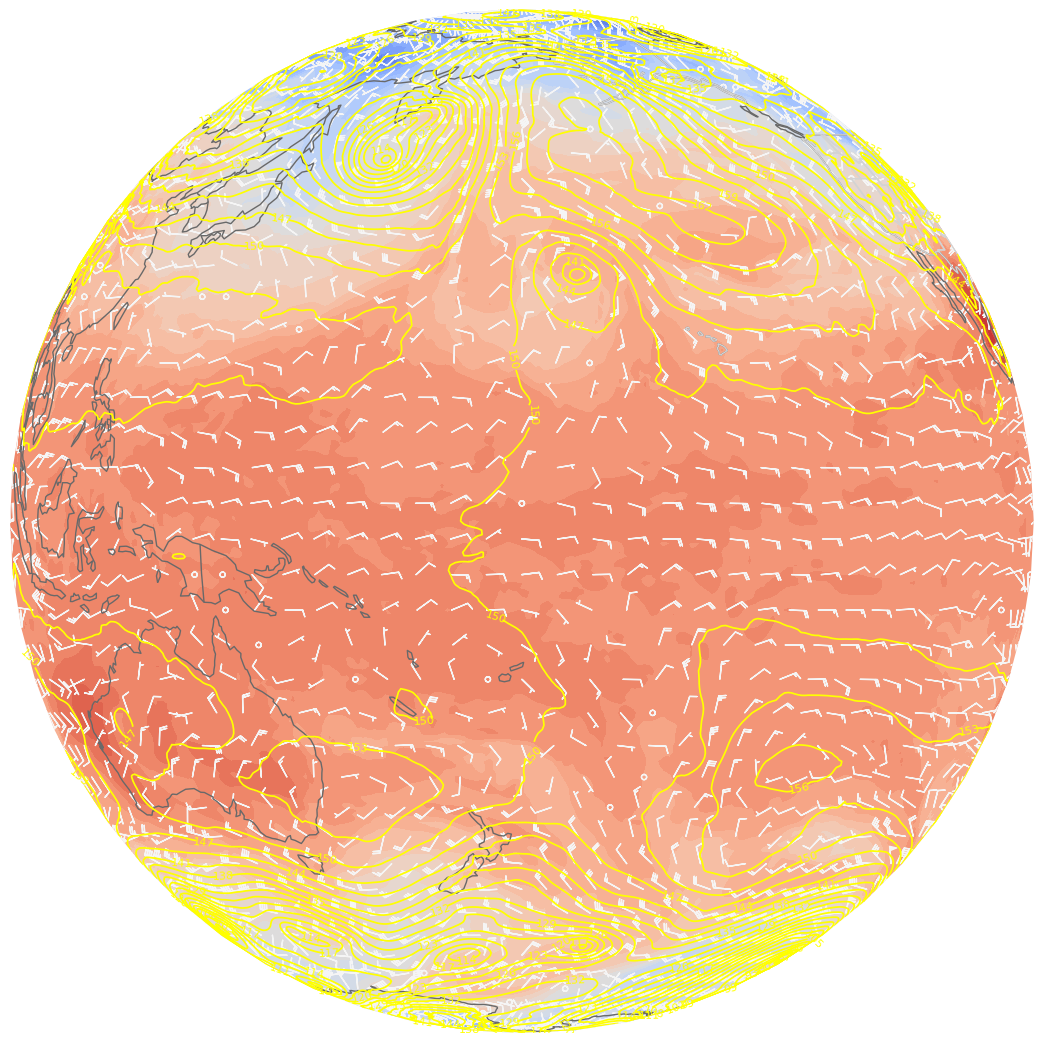
The seam has disappeared! Now we can go back to a PlateCarree projection, centered at the Greenwich Meridian (or wherever we’d like).
dpi = 100
fig = plt.figure(figsize=(2048/dpi, 1024/dpi))
ax = fig.add_subplot(1,1,1, projection=ccrs.PlateCarree(), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.BORDERS.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.STATES.with_scale(res), edgecolor=state_col, linewidth=0.75)
# Temperature (T) contour fills
# Note we don't need the transform argument since the map/data projections are the same, but we'll leave it in
CF = ax.contourf(lons_cyc,lats,T_cyc,levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
# Height (Z) contour lines
CL = ax.contour(lons_cyc,lats,Z_cyc,zContours,linewidths=1.25,colors='yellow', transform=proj_data)
ax.clabel(CL, inline_spacing=0.2, fontsize=8, fmt='%.0f')
fig.tight_layout(pad=.01)
# Plotting wind barbs uses the ax.barbs method. Here, you can't pass in the DataArray directly; you can only pass in the array's values.
# Also need to sample (skip) a selected # of points to keep the plot readable.
skip = 8
# Default length of wind barbs is 7. Shorten them a bit.
ax.barbs(lons_cyc[::skip],lats[::skip],U_cyc[::skip,::skip], V_cyc[::skip,::skip], color=wind_col, length = 5, transform=proj_data)
# Save this figure to your current directory
fig.savefig('test_ERA5_SOS.png')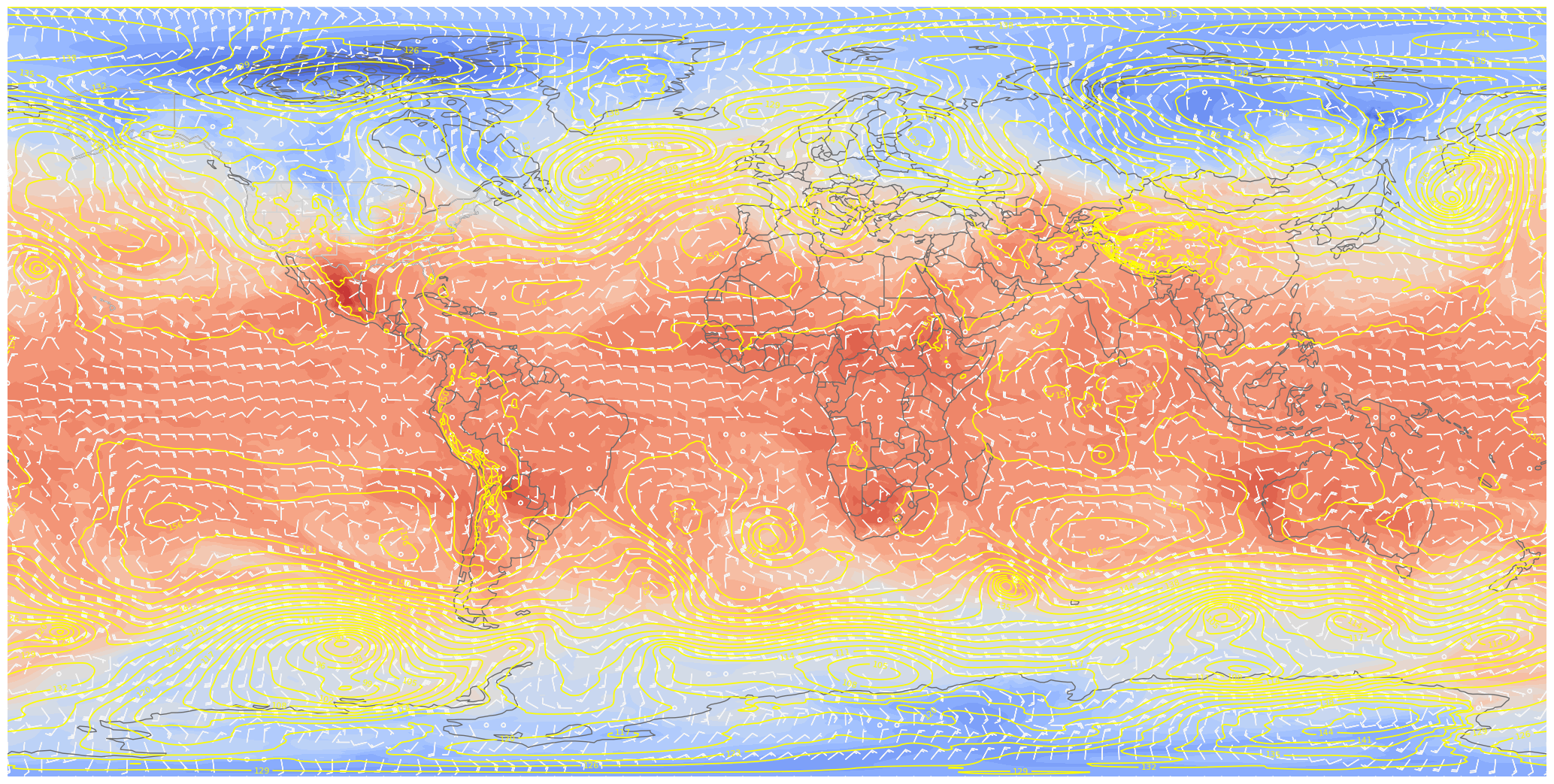
If you’d like, go to https://
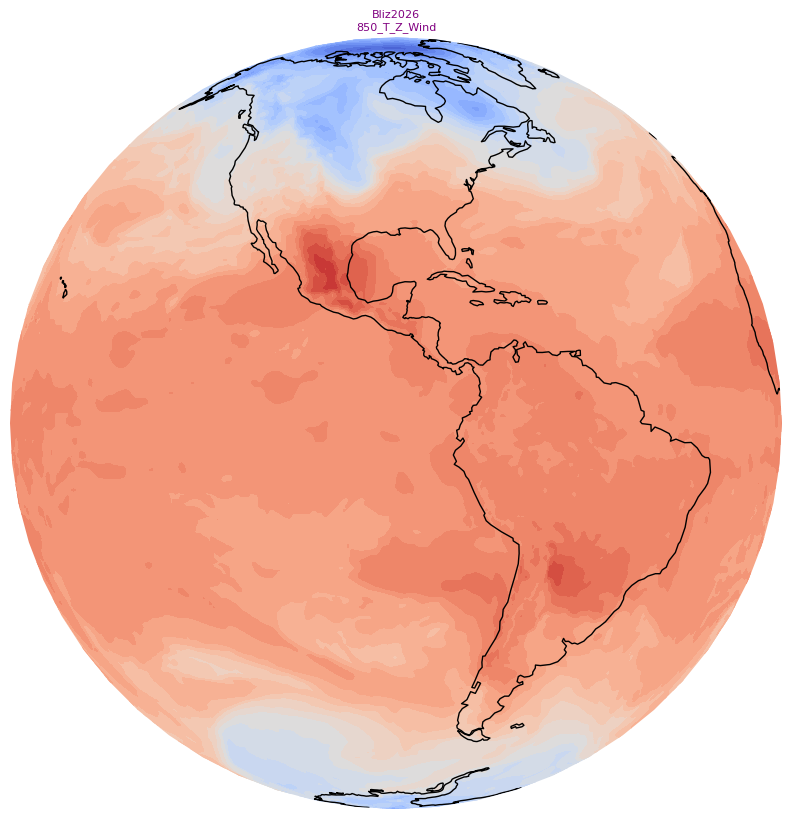
Create small (128x128) and large (800x800) thumbnails. These will serve as icons in the SoS iPad and computer apps that go along with your particular product.¶
We’ll use the orthographic projection and omit the contour lines and some of the cartographic features, and add a title string.
res = '110m'
dpi = 100
for size in (128, 800):
if (size == 128):
sizeStr = 'small'
else:
sizeStr ='big'
fig = plt.figure(figsize=(size/dpi, size/dpi))
ax = plt.subplot(1,1,1,projection=ccrs.Orthographic(central_longitude=-90), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res))
tl1 = caseDir
tl2 = prodDir
ax.set_title(f"{tl1}\n{tl2}", color='purple', fontsize=8)
# Contour fills
CF = ax.contourf(lons_cyc,lats,T_cyc, levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
fig.tight_layout(pad=.01)
fig.savefig(f'{mediaDir}thumbnail_{sizeStr}.jpg')
Create a standalone colorbar¶
Visualizations on the Science-on-a-Sphere consist of a series of image files, layered on top of each other. In this example, instead of having the colorbar associated with 850 hPa temperatures adjacent to the map, let’s use a technique by which we remove the contour plot, leaving only the colorbar to be exported as an image. We change the colorbar’s orientation to horizontal, and also change its tick label colors so they will be more visible on the sphere’s display.
# draw a new figure and replot the colorbar there
fig,ax = plt.subplots(figsize=(14,2), dpi=125)
# set tick and ticklabel color
tick_color='black'
label_color='orange'
cbar = fig.colorbar(CF, ax=ax, orientation='horizontal')
cbar.ax.xaxis.set_tick_params(color=tick_color, labelcolor=label_color, labelsize=8)
# Remove the Axes object ... essentially the contours and cartographic features ... from the figure.
ax.remove()
# All that remains is the colorbar ... save it to disk. Make the background transparent.
fig.savefig(f'{labelsDir}colorbar.png',transparent=True)
Create a set of labels for each plot¶
labelFname = f'{labelsDir}labels.txt'Define a function to create the plot for each hour.¶
The function accepts a time array element as its argument.
def make_sos_map (timeIdx):
res = '110m'
dpi = 100
fig = plt.figure(figsize=(2048/dpi, 1024/dpi))
ax = plt.subplot(1,1,1,projection=ccrs.PlateCarree(), frameon=False)
ax.set_global()
ax.add_feature(cfeature.COASTLINE.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.BORDERS.with_scale(res), edgecolor=coast_border_col)
ax.add_feature(cfeature.STATES.with_scale(res), edgecolor=state_col, linewidth=0.75 )
# add cyclic points to data and longitudes
# latitudes are unchanged in 1-dimension
Z_cyc, lons_cyc= cutil.add_cyclic_point(Zdam.isel(time=timeIdx), lons)
T_cyc, lons_cyc = cutil.add_cyclic_point(Tc.isel(time=timeIdx), lons)
U_cyc, lons_cyc = cutil.add_cyclic_point(Ukts.isel(time=timeIdx), lons)
V_cyc, lons_cyc = cutil.add_cyclic_point(Vkts.isel(time=timeIdx), lons)
# Temperature (T) contour fills
# Note we don't need the transform argument since the map/data projections are the same, but we'll leave it in
CF = ax.contourf(lons_cyc,lats,T_cyc,levels=tContours,cmap=plt.get_cmap('coolwarm'), extend='both', transform=proj_data)
# Height (Z) contour lines
CL = ax.contour(lons_cyc,lats,Z_cyc,zContours,linewidths=1.25,colors='yellow', transform=proj_data)
ax.clabel(CL, inline_spacing=0.2, fontsize=8, fmt='%.0f')
fig.tight_layout(pad=.01)
# Plotting wind barbs uses the ax.barbs method. Here, you can't pass in the DataArray directly; you can only pass in the array's values.
# Also need to sample (skip) a selected # of points to keep the plot readable.
skip = 8
# Default length of wind barbs is 7. Shorten them a bit.
ax.barbs(lons_cyc[::skip],lats[::skip],U_cyc[::skip,::skip], V_cyc[::skip,::skip], color=wind_col, length = 5, transform=proj_data)
frameNum = f'{timeIdx}'.zfill(2)
figName = f'{graphicsDir}ERA5_{caseDir}_{prodDir}_{frameNum}.png'
fig.savefig(figName)
# Reduce the size of the PNG image via the Linux pngquant utility. The -f option overwites the resulting file if it already exists.
# The output file will end in "-fs8.png"
! pngquant -f {figName}
# Remove the original PNG
! rm -f {figName}
# Do not show the graphic in the notebook
plt.close()Create the graphics and titles¶
Open a handle to the labels file
Define the time dimension index’s start, end, and increment values
Loop over the period of interest
Perform any necessary unit conversions
Create each timestep’s graphic
Write the title line to the text file.
Close the handle to the labels file
For demonstration purposes, we will use only produce graphics for the first two timesteps ... thus only 2 graphics will be produced. When you are ready, change the nFrames to nTimes so all times are processed.
nTimes = len(dateRange)
nTimes20labelFileObject = open(labelFname, 'w')
nFrames = 2
nFrames = nTimes
for timeIdx in range(0, nFrames, 1):
make_sos_map(timeIdx)
# Construct the title string and write it to the file
valTime = pd.to_datetime(times[timeIdx].values).strftime("%m/%d/%Y %H UTC")
tl1 = f'ERA5 {p_level} hPa Z/T/Wind {valTime} \n' # \n is the newline character
labelFileObject.write(tl1)
print(tl1)
# Close the text file
labelFileObject.close()
print("Finished with SoS graphic loop")ERA5 850 hPa Z/T/Wind 02/20/2026 00 UTC
ERA5 850 hPa Z/T/Wind 02/20/2026 06 UTC
ERA5 850 hPa Z/T/Wind 02/20/2026 12 UTC
ERA5 850 hPa Z/T/Wind 02/20/2026 18 UTC
ERA5 850 hPa Z/T/Wind 02/21/2026 00 UTC
ERA5 850 hPa Z/T/Wind 02/21/2026 06 UTC
ERA5 850 hPa Z/T/Wind 02/21/2026 12 UTC
ERA5 850 hPa Z/T/Wind 02/21/2026 18 UTC
ERA5 850 hPa Z/T/Wind 02/22/2026 00 UTC
ERA5 850 hPa Z/T/Wind 02/22/2026 06 UTC
ERA5 850 hPa Z/T/Wind 02/22/2026 12 UTC
ERA5 850 hPa Z/T/Wind 02/22/2026 18 UTC
ERA5 850 hPa Z/T/Wind 02/23/2026 00 UTC
ERA5 850 hPa Z/T/Wind 02/23/2026 06 UTC
ERA5 850 hPa Z/T/Wind 02/23/2026 12 UTC
ERA5 850 hPa Z/T/Wind 02/23/2026 18 UTC
ERA5 850 hPa Z/T/Wind 02/24/2026 00 UTC
ERA5 850 hPa Z/T/Wind 02/24/2026 06 UTC
ERA5 850 hPa Z/T/Wind 02/24/2026 12 UTC
ERA5 850 hPa Z/T/Wind 02/24/2026 18 UTC
Finished with SoS graphic loop
Create an SoS playlist file¶
We follow the guidelines in https://
playlistFname = f'{playlistDir}playlist.sos'Open a file handle to the playlist file
Add each line of the playlist. Modify as needed for your case and product; in general you will only need to modify the name and description values!
Close the file handle
# Set the author (creator) name
creaName = "Rovin Lazyle" # Put your name here!
plFileObject = open(playlistFname, 'w')
subCat = "ERA5"
cRet = "\n" # New line character code
lBr = "{"
rBr = "}"
nameStr = f'ERA5 {caseDir}{prodDir}'
descriptionStr = f'{lBr}{caseDir}{prodDir}{rBr}'
creaName = "Rovin Lazyle" # Put your name here!
plFileObject.write(f"name = {nameStr}{cRet}")
plFileObject.write(f"description = {descriptionStr}{cRet}")
plFileObject.write(f"pip = ../labels/colorbar.png{cRet}")
plFileObject.write(f"pipheight = 10{cRet}")
plFileObject.write(f"pipvertical = -35{cRet}")
plFileObject.write(f"label = ../labels/labels.txt{cRet}")
plFileObject.write(f"layer = Grids{cRet}")
plFileObject.write(f"layerdata = ../2048{cRet}")
plFileObject.write(f"firstdwell = 2000{cRet}")
plFileObject.write(f"lastdwell = 3000{cRet}")
plFileObject.write(f"fps = 8{cRet}")
plFileObject.write(f"zrotationenable = 1{cRet}")
plFileObject.write(f"zfps = 30{cRet}")
plFileObject.write(f"subcategory = {subCat}{cRet}")
plFileObject.write(f"Creator = {creaName}{cRet}")
plFileObject.close()Display the contents of the playlist file
! cat {playlistFname}name = ERA5 Bliz2026850_T_Z_Wind
description = {Bliz2026850_T_Z_Wind}
pip = ../labels/colorbar.png
pipheight = 10
pipvertical = -35
label = ../labels/labels.txt
layer = Grids
layerdata = ../2048
firstdwell = 2000
lastdwell = 3000
fps = 8
zrotationenable = 1
zfps = 30
subcategory = ERA5
Creator = Rovin Lazyle
The playlist essentially describes the following:
The name and description of the product, which are used in the SoS iPad and computer apps
The three components of what is displayed on the screen:
The colorbar (a picture-in-picture, aka pip), with its height and vertical position
The label, or title, which describes the graphic
The graphic layer itself. Multiple graphic layers, each with its own layer name and directory path, could be included; in this case, there is just one.
The dwell times, in ms, for the first frame, last frame, and all frames in-between.
We’re done! The directory tree you have created can then be copied/synced to the correct directory on the SoS computer (Ross and Kevin will take care of this).¶
Summary¶
We now have an end-to-end workflow that will create all that is necessary for a custom SoS visualization, using ERA5 reanalysis data.
Things to try¶
First, simply pick a 2-7 day date range of interest to you ... perhaps for your case study. Re-run the notebook.
Next, try different levels and variables.
After that, try adding a gridded diagnostic to your global maps.
What’s Next?¶
We will streamline this notebook, so it can be used as a template for your own case.